All-cause Mortality risk assessment
Our health is an intricate interplay between our genes, our epigenetics, and our environment. However perplexing these interactions may seem, we are getting closer to be able to put an accurate number on the risk of mortality. Much more so, than our previous guesswork based on chronological age and family histories.
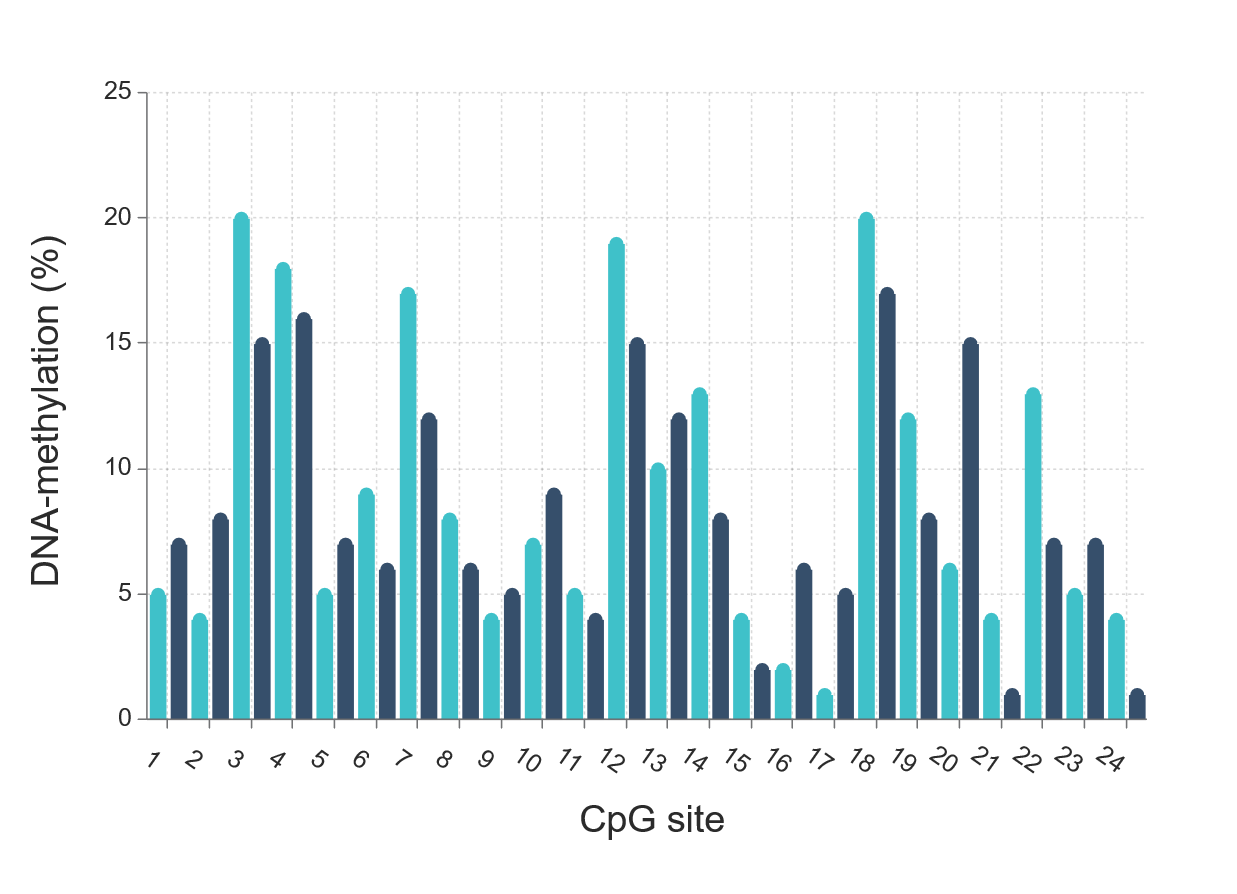
As well as estimating biological age, the epigenetic clock has also been shown to predict all-cause mortality (See Biological Age Determination). Many of the DNA methylation sites in the genome (CpGs) which are used in the epigenetic clock however, are not directly related to genes implicated in mortality themselves. Rather, when the individual CpGs are read together, they provide a comprehensive readout of the epigenome determining biological age; illustrating that the whole is greater than the sum of its parts. Newer versions of the clock (such as DNAm PhenoAge) incorporate more sites which are directly linked to diseases and indeed outperform previous versions in predicting mortality risk.

It seems logical, that in order to improve our risk estimates we have to study CpGs that are directly linked to mortality and disease. A recent study identified 58 CpGs that strongly predicted all-cause mortality over a 14 year follow-up period, independently of the sites used in the calculation of the epigenetic clock.
The CpGs were associated with genes implicated in coronary artery disease, Parkinson’s disease, neuropsychiatric disorders, multiple sclerosis, and different types of cancers such as lung, brain, colon, breast, skin, and pancreatic cancer.
Interestingly, the methylation patterns of many implicated CpGs in the study were strongly associated with smoking, reinforcing the importance of lifestyle decisions as risk factors. Our models at EPIGEN are based on the latest research and are able to predict all-cause mortality with high accuracy.